38 product results for:
"GFP"
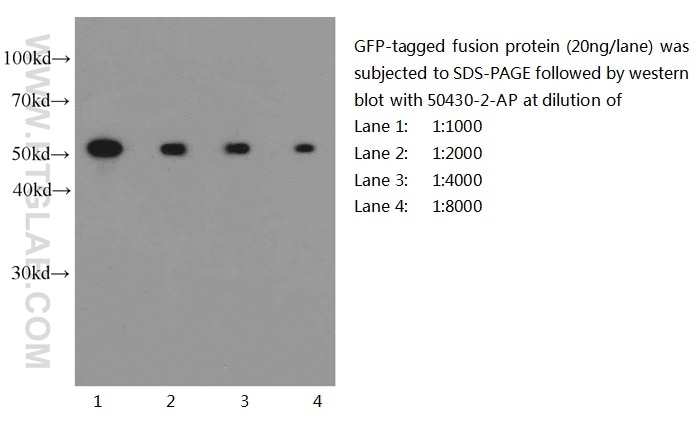
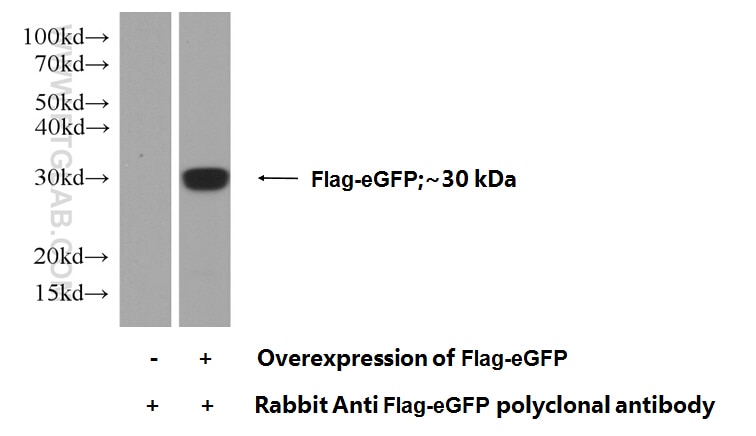
GFP primary antibodies (13)
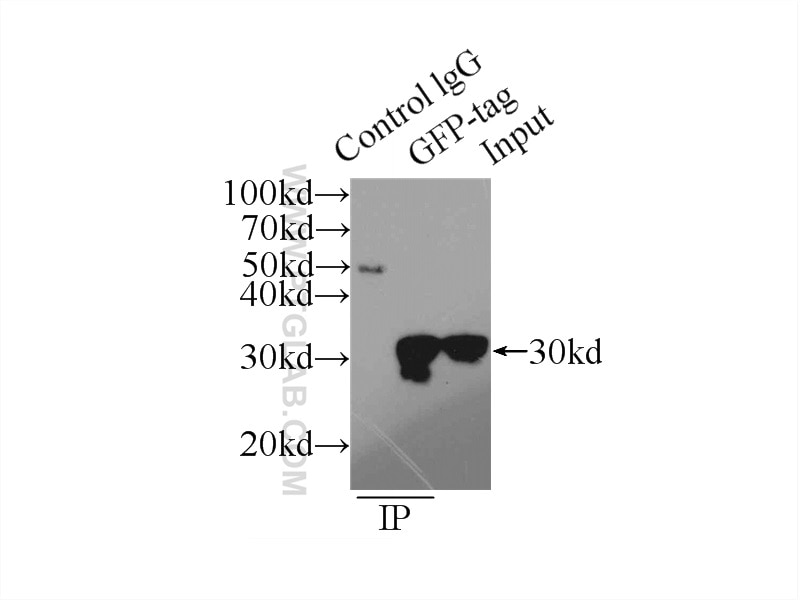
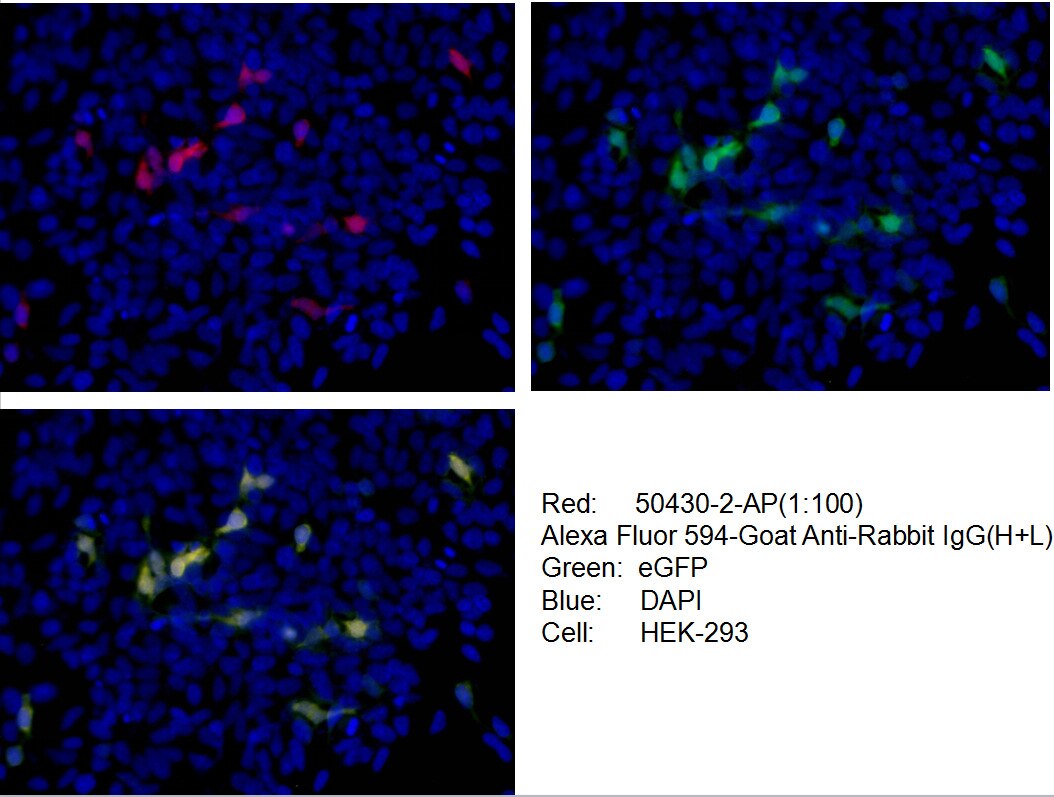
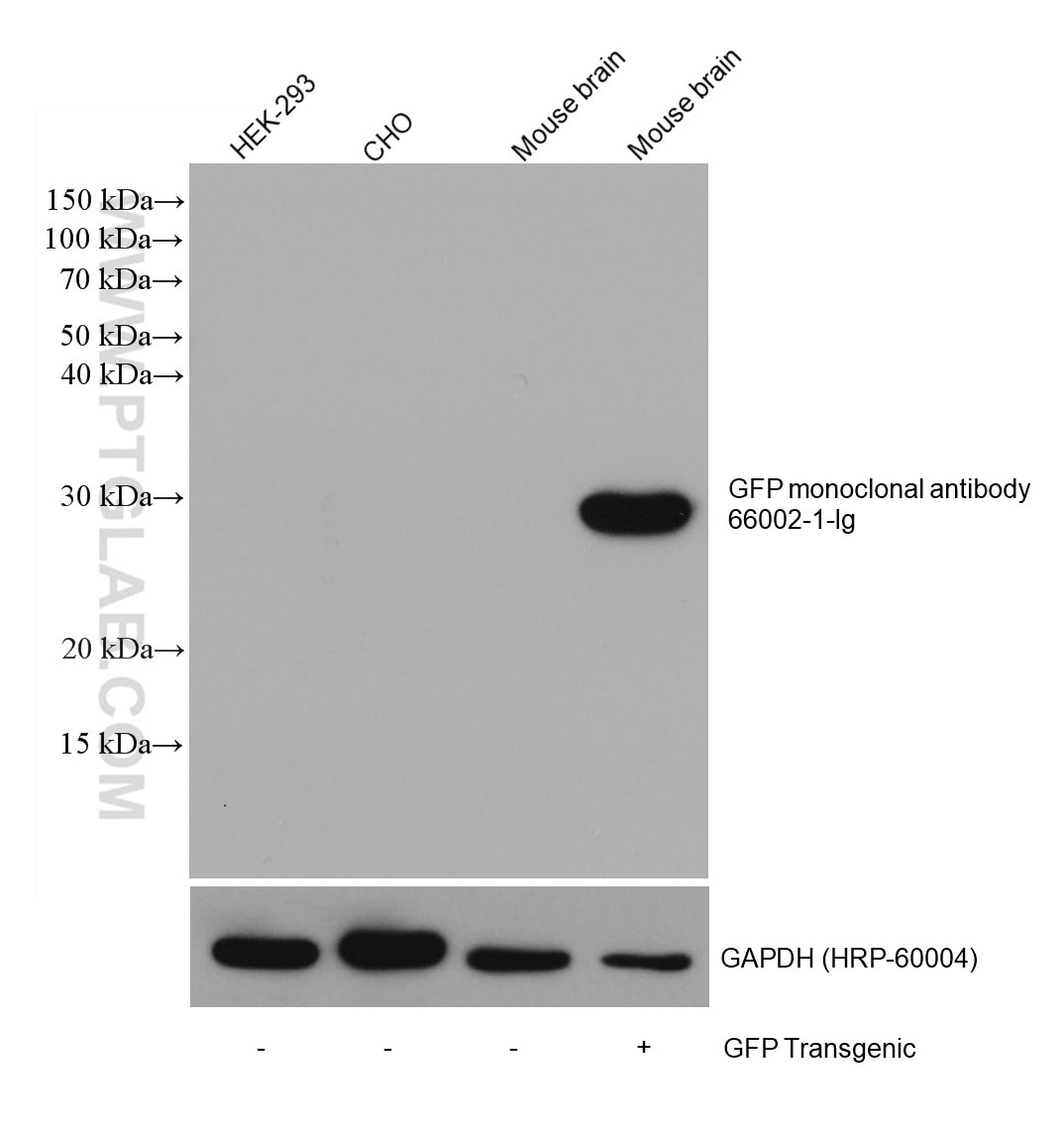
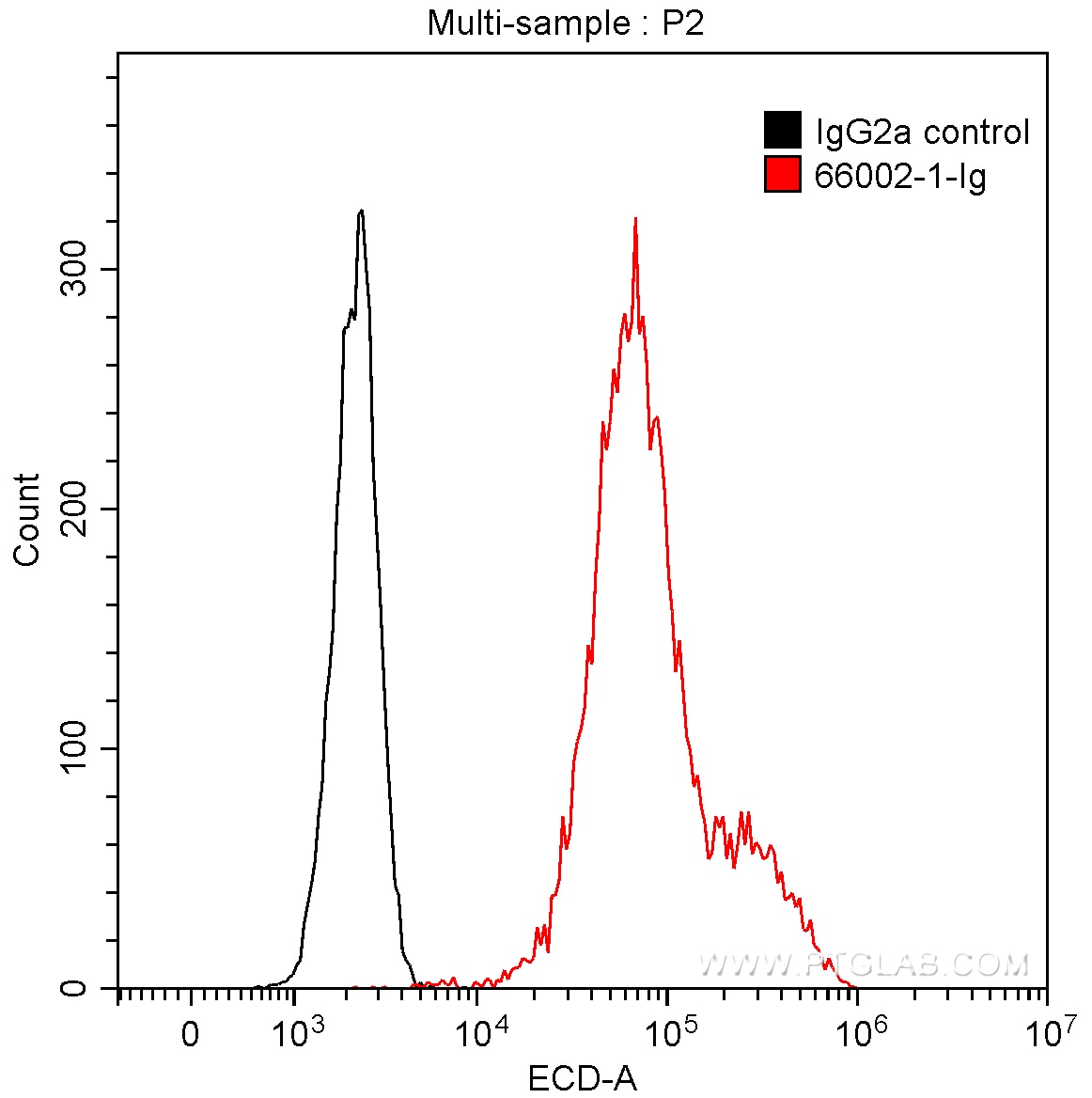
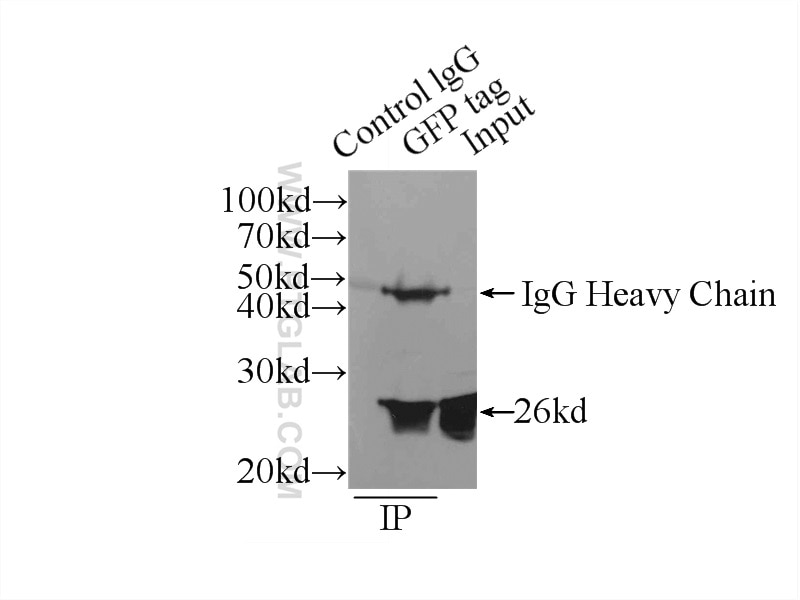
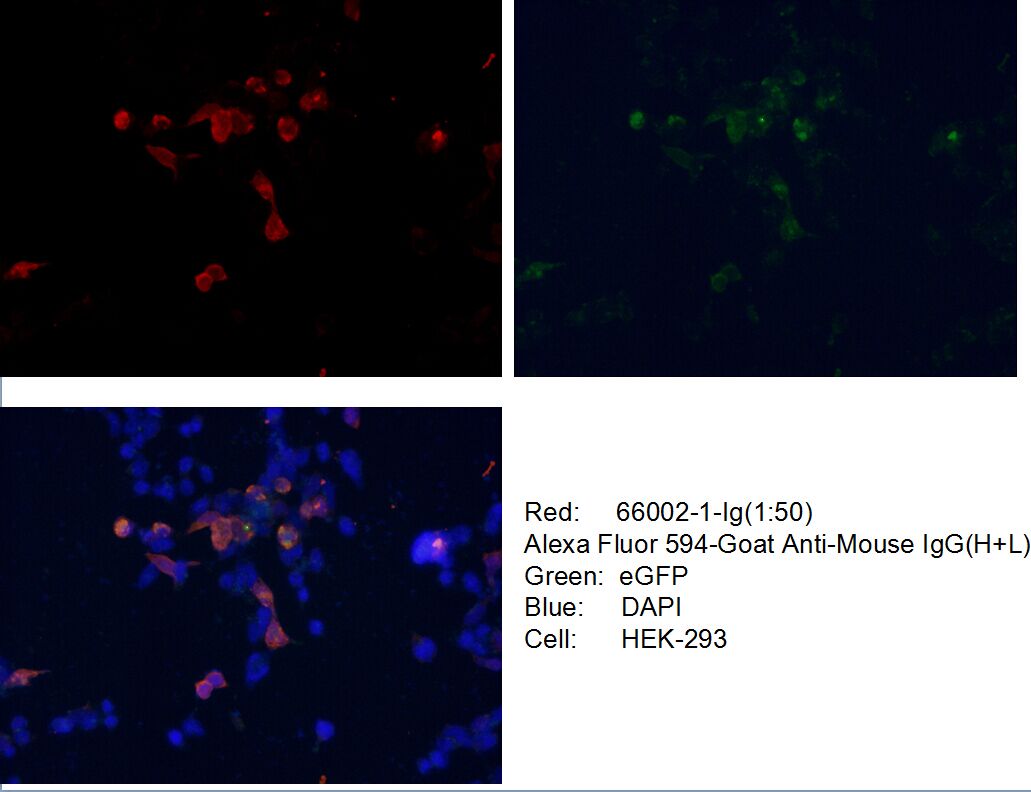
Conjugated and unconjugated polyclonal/monoclonal GFP antibodies available. Validated in WB, IP, IHC, IF, FC, CoIP, ChIP, RIP, IP-MS and ELISA.
GFP nanobodies (22)
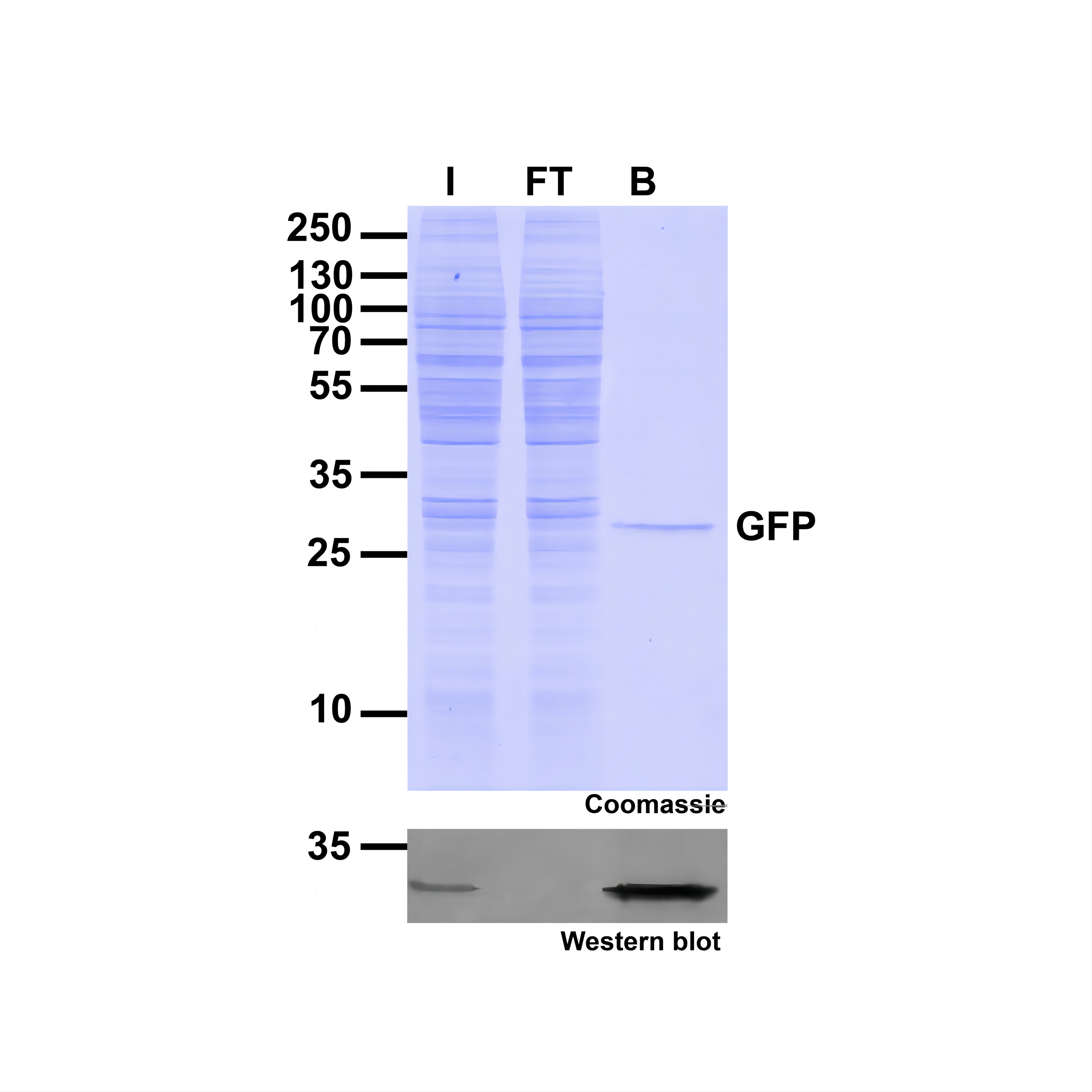
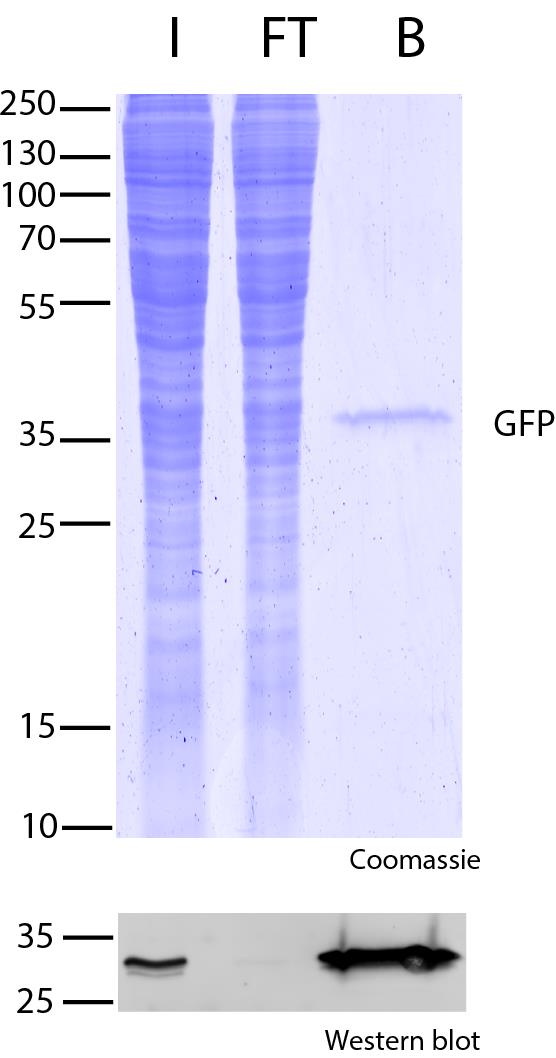
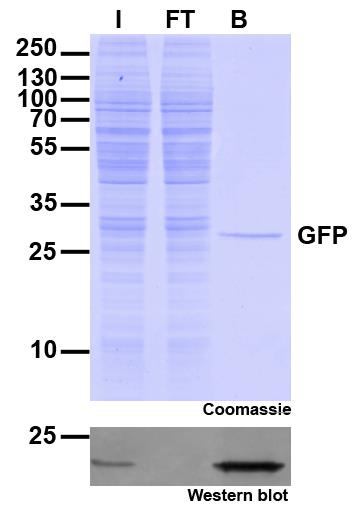
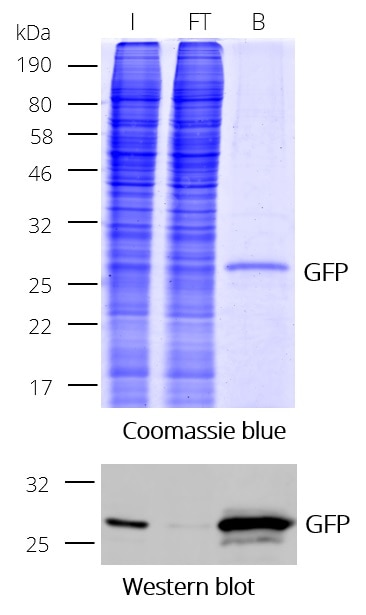
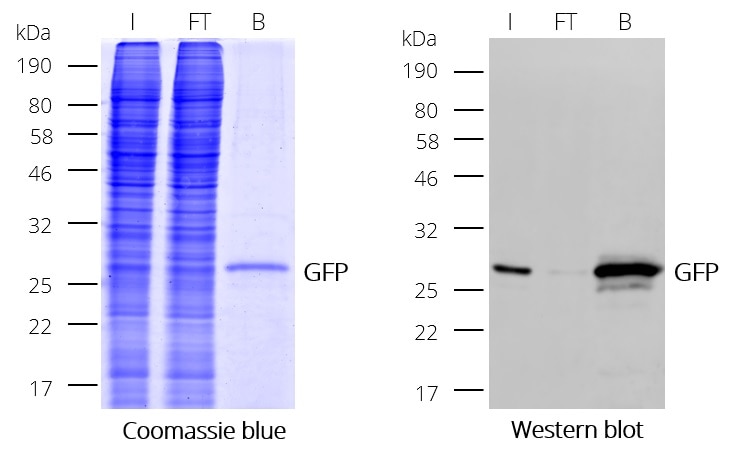
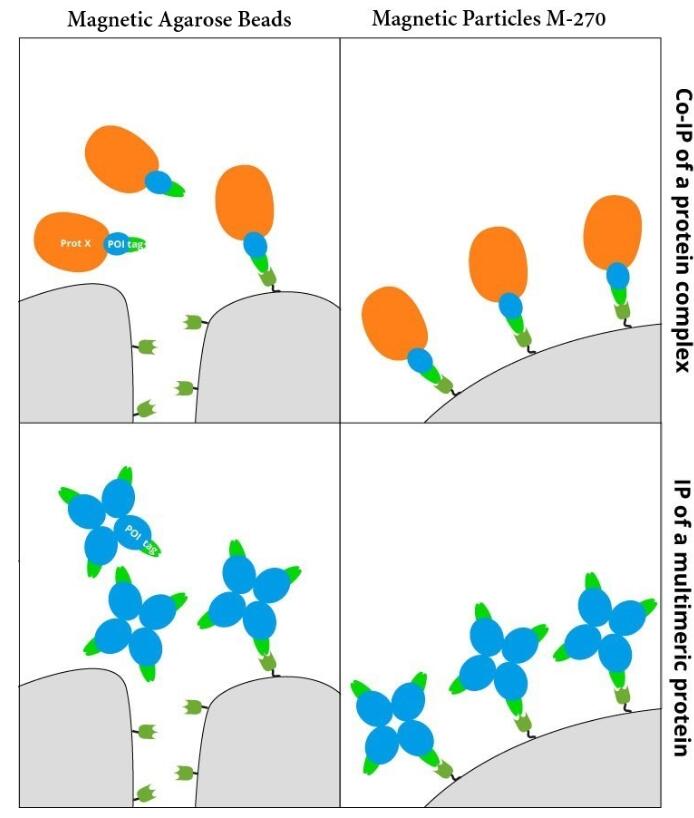
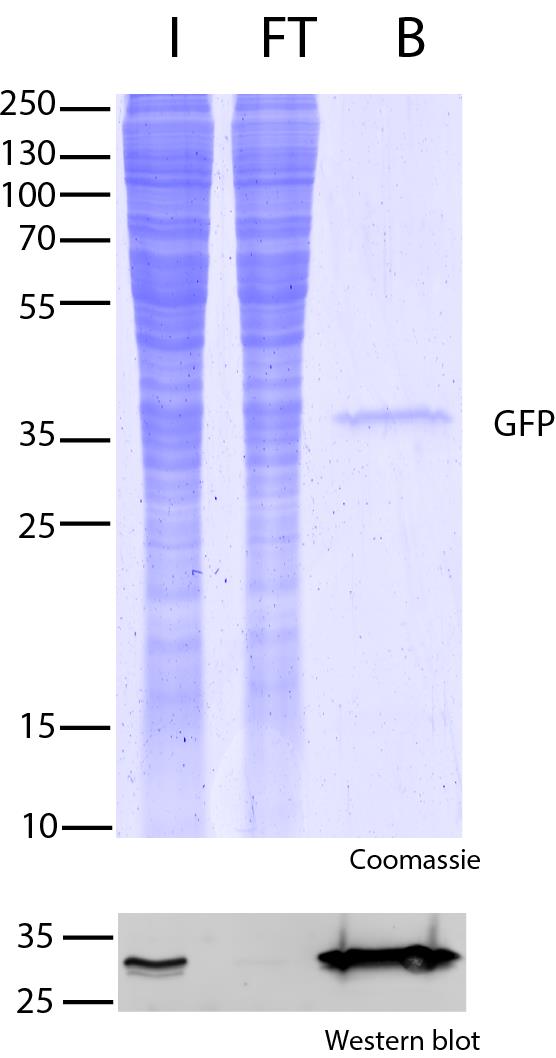
Ready-to-use affinity resin for immunoprecipitation (IP) of GFP-fusion proteins. The GFP-Trap® is an anti-GFP Nanobody/ VHH coupled to agarose beads, magnetic agarose beads, magnetic particles or multiwell plates. Tester for IP, CoIP, MS, RIP and enzyme activity measurements. Learn more about nanobodies.
GFP Recombinant (3)
E. coli expressed GFP tag fusion proteins.