
Mass Spectrometry
Mass spectrometry (MS) is a common method for identification of proteins. MS is particularly helpful for protein-protein interaction assays or interactome studies, which aim to identify the interacting partners of a protein within a cell at a time point.
Nano-Traps are beneficial for MS analysis following Co-IP, because of their reliable performance, high affinity, and low background of protein contaminations. The superior stability of the Nano-Traps allows to apply stringent washing conditions. In most cases, even buffers containing chaotropic reagents can be used, whereas these would inactivate other IgG-based or Streptavidin-based affinity resins. Therefore, background can be further reduced, which increases the sensitivity of the sample analysis. This may be required for analysis of some post-translational modifications (PTMs).
It is recommended to conduct the MS sample preparation on-bead to avoid loss of sample by incomplete elution from the Nano-Trap.
Immunoprecipitation mass spectrometry (IP-MS) is also called affinity purification mass spectrometry (AP-MS) or affinity capture mass spectrometry (affinity capture MS).
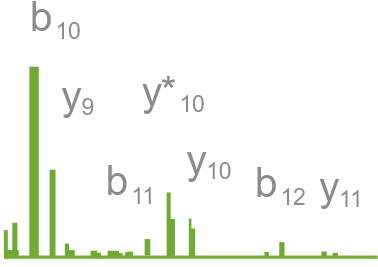
Section of mass spectrum